Especially in the light of climate change, we need fundamentally new concepts to generate and use environmentally friendly and climate-compatible forms of energy, so-called 'green energy'. The main focus of the Photobiotechnology group is the study and biotechnological application of hydrogen-forming enzymes (hydrogenases) and other redox enzymes, which assume important functions in microbial energy metabolism (we mainly study photosynthetic microorganisms) or which catalyze industrially relevant chemical syntheses. Our future-oriented, internationally connected research with strong links to industrial biotechnology is conducted at the interface of microbiology, biochemistry and chemical biology. To elucidate structure-function relationships of biotechnologically relevant enzymes, we use a variety of state-of-the-art genetic, biochemical and biophysical methods and cooperate nationally and internationally with leading scientists (e.g. Oxford, Osaka etc). Applying enzyme engineering, we also create bio-inspired novel biocatalysts which can be used in a future application in the context of a bio-based industry as miniaturized, robust catalysts in H2 technologies and in 'green' chemistry.
In addition to our focus on protein research, we characterize stress responses of microalgae (also see Hemschemeier). In view of climate change, extreme weather conditions are expected to occur frequently in the future. How plants and algae adapt to these is of crucial importance for the Earth's ecosystem and our climate. We investigate the acclimation of microalgae to stress conditions, especially hypoxia, on the physiological, genetic and biochemical level. One focus is the study of stress-induced nitric oxide-based signal transduction. Here we characterize heme-binding proteins (microbial hemoglobins and NO-sensitive guanylate cyclases) biochemically and biophysically, especially with regard to NO-relevant reactions. Furthermore, we investigate signal transduction components and regulated target proteins in physiological terms by identifying interaction partners and generating mutants of the corresponding genes using CRISPR/Cas technology. Since some microalgae produce hydrogen in response to stress, there is complementary overlap and interdisciplinary collaboration within the group as well as in our international collaborative network.
Hydrogenases catalyze the reversible reduction of protons to molecular hydrogen (H2). They play an important role in the energy metabolism of many prokaryotic and eukaryotic microorganisms, but are also of great interest for the biobased production and utilization of hydrogen. Evolution has produced different classes of hydrogenases that are distinguished by their active sites. We explore the particularly efficient so-called [FeFe]-hydrogenases, which contain an active site unique in nature, the H-cluster: Coupled to a 'classical' [4Fe4S] cluster via a coordinating cysteine residue is a unit consisting of two iron ions coordinated by the biologically unusual ligands cyanide and carbon monoxide as well as an azadithiolate group (see figure). It is at this unit that the actual catalysis takes place. We are trying to understand the molecular mechanisms that make [FeFe]-hydrogenases such efficient biocatalysts. For example, we use high-resolution crystal structures and modern spectroscopic techniques such as infrared or electron spin resonance spectroscopy to investigate how the protein scaffold and the inorganic H-cluster interact. To this aim, we also produce variants of [FeFe] hydrogenases using molecular biology methods (site directed mutagenesis) to investigate the influence of individual amino acid residues. For example, we were able to help unravel the conserved and highly efficient proton transport pathway in the protein molecule.
We are also interested in how the H-cluster is assembled in the cell. According to current knowledge, the [4Fe4S] cluster is produced by the housekeeping FeS cluster assembly machinery, whereas production of the two-iron moiety requires three special maturation proteins found exclusively in [FeFe]-hydrogenase-containing organisms. These maturation factors catalyze complex reactions to produce the unusual active site from amino acids and iron and subsequently transfer it to the hydrogenase protein.
In living cells – but also for industrial applications – hydrogenases must interact with redox partners that provide or accept electrons. In nature, these are often small redox proteins such as ferredoxins, cytochromes or flavodoxins. We are interested in the molecular interactions of the proteins and how electron transport pathways are coordinated in the cell. Of particular interest are the [FeFe]-hydrogenases in the chloroplasts of unicellular green algae such as the model organism Chlamydomonas reinhardtii. These hydrogenases interact with photosynthetic ferredoxin and, under certain conditions, obtain electrons from the photosynthetic electron transport chain. Hence, the light-driven provision of high-energy redox equivalents is coupled with the production of H2, whereas in non-photosynthetic organisms or industrial processes, high-energy substrates or electricity must be provided. While this metabolic pathway is of great interest for basic research and ecology, we would also like to use it for applied processes. To this end, we are trying to optimize the H2-generating processes in the algae, but also to combine and optimize the proteins of these metabolic pathways in the test tube.
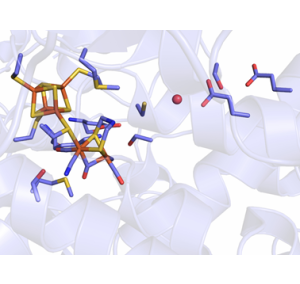
[FeFe]-hydrogenases are extremely efficient and have the advantage over platinum-based catalysts that they use iron, which is found in large quantities on earth. However, it has not yet been possible to integrate them into economically relevant processes. One reason for this is their pronounced sensitivity to molecular oxygen (O2), which irreversibly destroys the active site, the H-cluster, of almost all known [FeFe]-hydrogenases. One focus of our work is to understand this O2-sensitivity and to develop more O2-tolerant hydrogenases by means of random (directed evolution) as well as rational modifications to the protein. Several years ago, a naturally O2-stable [FeFe]-hydrogenase was discovered. We were able to structurally and biochemically characterize this novel hydrogenase and mechanistically explain the oxygen stability of the enzyme. A research topic of our group is now to identify and investigate further O2-stable [FeFe]-hydrogenases in order to design enzymes that can be stably integrated into applied processes.
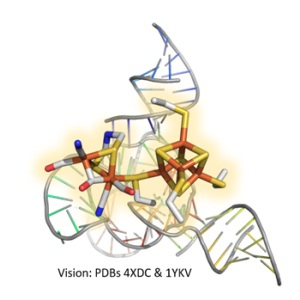
Furthermore, we are working on the development of small, robust minimal hydrogenases. Here we are collaborating with chemists who can synthesize the two-iron active site unit described in "Molecular mechanisms of hydrogenases and their redox partners". Indeed, we also use this procedure for the production of large amounts of hydrogenase proteins in Escherichia coli, the popular host for heterologous expression. E. coli does not have [FeFe]-hydrogenases itself and therefore cannot assemble the active site. However, we can convert hydrogenases produced in the bacterium to active holoprotein using the chemically synthesized two-iron moiety. This surprisingly simple procedure, which we have established, also allows us to work with chemists to incorporate modified cofactors into the hydrogenase protein in order to understand which molecular interactions produce an active biocatalyst for H2 formation. With this knowledge, we are now attempting to minimize the natural protein scaffold. On the one hand, we are investigating whether the H-cluster can be bound and activated by minimal peptides. Another, seemingly futuristic, stragety is the use of DNA building blocks to functionally integrate the H-cluster. DNA nanotechnology has made enormous progress in recent years and, in addition to micromaterials (e.g. DNA origami), can also provide redox-active nucleic acid scaffolds. Here we cooperate with scientists who are experienced in the synthesis and manipulation of DNA nanostructures.
We are convinced that living organisms can help us to establish sustainable industrial processes and to protect and regenerate our ecosystems. Organisms that perform oxygenic photosynthesis are particularly promising in this regard. These organisms draw the energy necessary for life from sunlight and require only inorganic molecules – such as CO2, nitrate or sulfate – to build up the complex living matter. In addition to our projects aimed at optimizing 'green' H2 formation by algae or by isolated enzymes from algae, we want to use additional elaborate biosynthetic pathways from photosynthetic microorganisms for biobased synthesis processes. So-called Old Yellow Enzymes provide one example. These flavin-containing and thus yellow-colored enzymes are en-reductases that catalyze a variety of redox reactions. Outside the living cell, they can produce many molecules for which there is already an industrial market, such as fragrances or insect-repellent substances. These enzymes also exist in algae and we are characterizing the proteins to potentially discover new or optimized properties. These enzymes also require a source of electrons, namely the well-known redox equivalent NADPH. Possibly of greater interest, therefore, is that we have discovered that living algal cells can also convert the relevant substances, and that this biotransformation depends on photosynthesis. Therefore, we are exploring the function and metabolic integration of Old Yellow enzymes in algae. In addition to the clear perspective for a biotechnological application, we are interested in the physiological function of these enzymes in algae, because almost nothing is known about this so far.
see research topic Hemschemeier
